Nana Matoba, Bran Le, and Jordan Valone released a preprint describing context dependent genetic variant effects. Gene regulatory effects in bulk-post mortem brain tissues are undetected at many non-coding brain trait-associated loci. We hypothesized that context-specific genetic variant function during stimulation of a developmental signaling pathway would explain additional regulatory mechanisms. We measured chromatin accessibility and gene expression following activation of the canonical Wnt pathway in primary human neural progenitors from 82 donors. Context-specific molecular quantitative trait loci increased brain-trait colocalizations by up to 70%, suggesting that genetic variant effects during early neurodevelopmental patterning lead to differences in adult brain and behavioral traits. The study can be found here.
Cell type specific genetic effects on post transcriptional modifications
Nil Aygün led a study on identifying genetic variants affecting RNA editing and alternative polyadenylation sites in human neural progenitors and their differentiated neuronal progeny. More RNA-editing and isoforms utilizing longer polyadenylation sequences were observed in neurons, likely due to higher expression of genes encoding the proteins mediating these post-transcriptional events. We also detected hundreds of cell-type-specific editing quantitative trait loci (edQTLs) and alternative polyadenylation QTLs (apaQTLs). We found colocalizations of a neuron edQTL in CCDC88A with educational attainment and a progenitor apaQTL in EP300 with schizophrenia, suggesting genetically mediated post-transcriptional regulation during brain development lead to differences in brain function. The study can be found here.
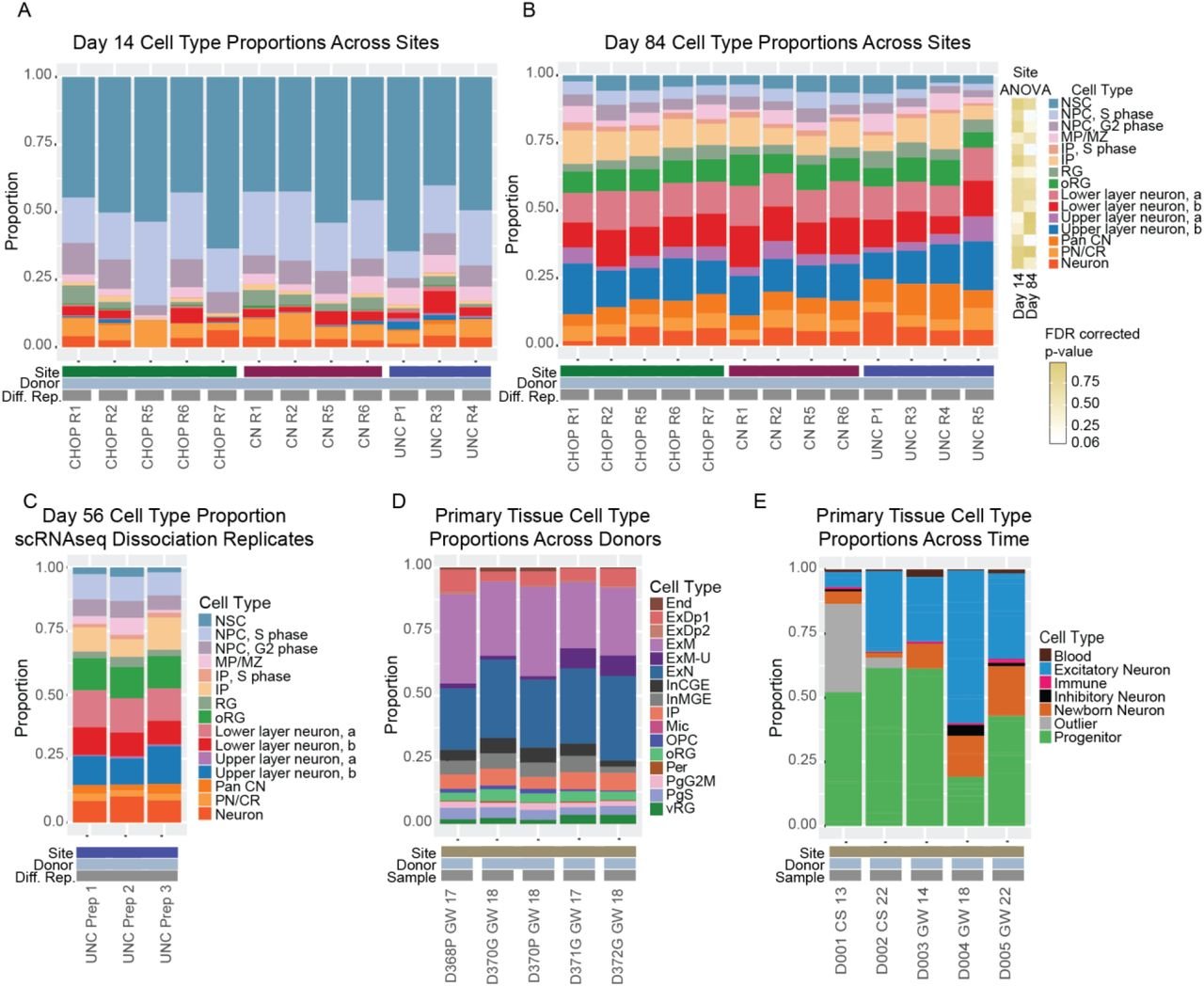
Cross-site reproducibility of cortical organoid differentiation
Rose Glass together with Elisa Waxman and Deb French at CHOP, and Satoshi Yamashita and Kazue Hashimoto-Torii at Children’s National released a preprint evaluating the reproducibility or cortical organoid differentiation. Cell type proportions were found to be largely reproducible across sites, but differences in gene expression were identified. The study can be found here.
Brandon Le successfully defends his dissertation
Brandon Le successfully defended his dissertation on stimulus specific genetic effects in human neural progenitor cells. He is now finishing up this work as a postdoc in the Stein lab. Congrats Bran!
Dr. Rose Glass successfully defends her dissertation
Rose successfully defended her dissertation on evaluating the reproducibility of human cortical organoids as well as using organoids to model brain overgrowth in autism. She is now a postdoc with Mustafa Sahin at Boston Children’s Hospital. Congrats Rose!
New diSPIM Microscope
The diSPIM has arrived! Great resolution and it’s fast. It’s beautiful!
Say Hello to Two New Graduate Students
We’re happy to welcome Miguel Cuevas and Rubal Singla to the lab. Both Miguel and Rubal will be working with brain organoids. Welcome to the team!
Stein Lab Welcomes a New Post-Doc
We welcome Jieun (Esther) to the team. She will be working with brain organoids during her time at the lab. Welcome Esther!
Nana Promoted to Research Associate
Effective January 3, 2023, Nana is promoted to research associate at the Stein Lab. Thank you for being an integral part of the lab!
Nil Aygün Successfully Defends Her Dissertation
Dr. Nil Aygün successfully defended her dissertation in March 2023, discussing the genetics of cell-type specific gene regulation and drug-sensitive cellular phenotypes during human neurogenesis.
Talk on Drug Discovery News
Stein Lab Holiday Party 2022
Grateful to work with all these brilliant scientists. Happy holidays from the Stein Lab!
Veda Dayananda places at Science Competitions
High school student Veda Dayananda placed 2nd at the NC Student Academy of Science State competition (https://www.ncsas.org/) and placed 3rd at the NC State Science & Engineering Fair (http://ncsef.org/) for her project entitled “Quantifying Region-Specific Cortical Volumes in a Mouse Modeling an Autism-Associated Mutation”. Congrats, Veda!
Justin Wolter wins K01 award
Justin Wolter won a K01 award to support his transition to faculty. The award is entitled “Discovery of genetic modifiers of PTEN-ASD severity in a library of genetically diverse iPSC lines”. Congrats, Justin!
Genetic variants modulating proliferative response to lithium
In a study led by Justin Wolter and Bran Le, we found and validated genetic variants altering lithium’s proliferative response in neural progenitor cells. This study opens up a new research direction for our lab, cellular pharmacogenomics. This study was published as a Priority Communication in Biological Psychiatry and was featured in an Elsevier press release.
miRNA eQTLs in the developing cortex
Mike Lafferty led a study to find how genetic variants influence miRNAs in the developing human cortex. Genetic variants associated with one miRNA (miR4707) were also associated with brain size and educational attainment. We suggest that this miRNA has important and previously unknown functions during human neurogenesis. The pre-print describing these findings is found here.
Inferring cell-type-specific causal gene regulatory networks during human neurogenesis
Nil Aygün led a study using causal inference tools to infer how genetic variants influence gene expression through changes in chromatin accessibility and through changes in expression of other genes. This allows an understanding of how genetic variation leads to changes in gene expression within specific cell types present during neurogenesis. This study is available as a pre-print here.
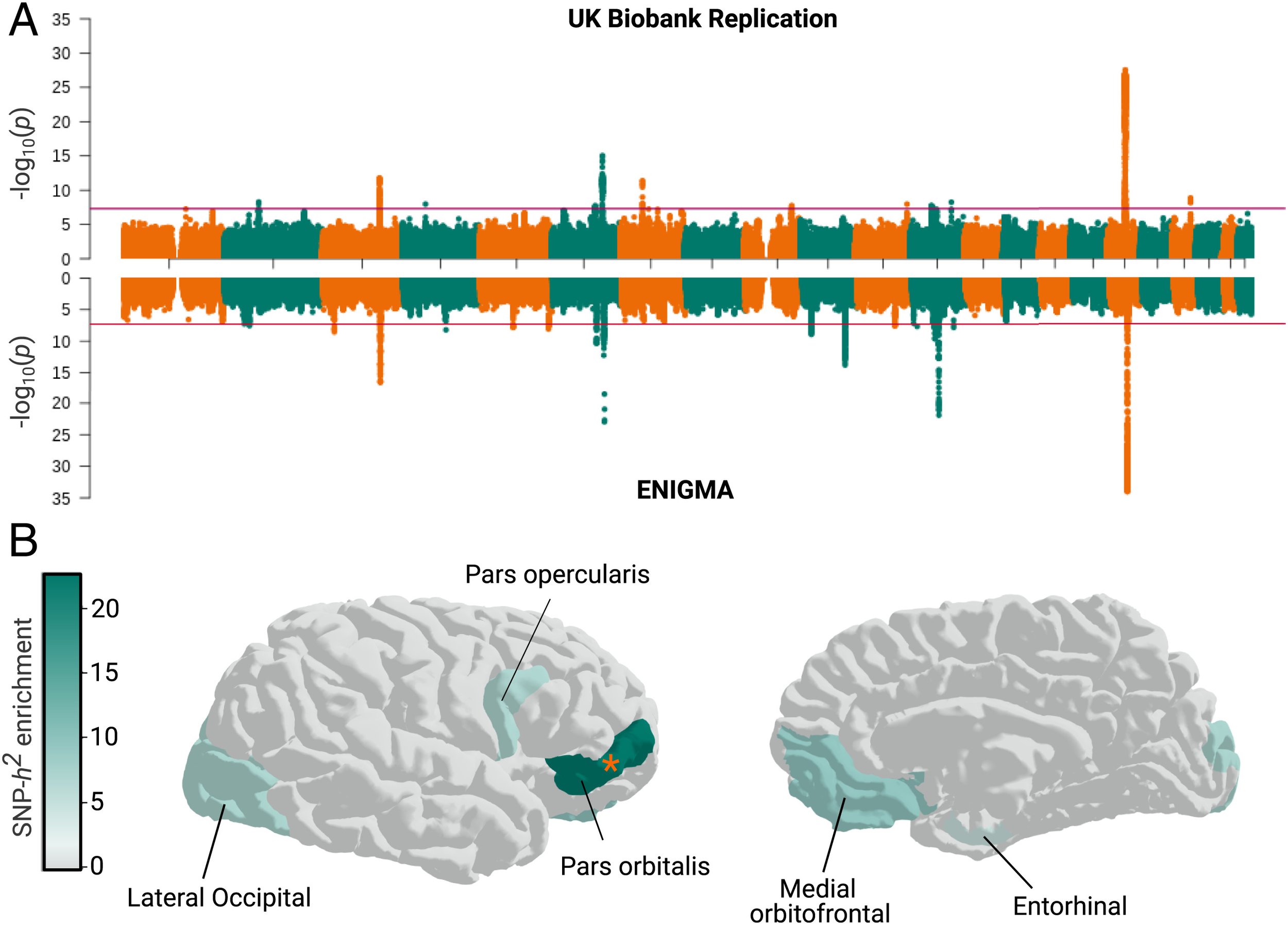
Using genetics to understand how evolution shaped modern human brain structure
Led by the lab of Simon Fisher, we attempted to replicate some of our previous findings on how evolution has changed the size and shape of the human brain. We found that our previous evidence of recent selection influencing cortical structure was not replicated in an independent sample. But, human gained enhancers did have a replicable effect on human brain structure. This study was published in PNAS.
MPRA review published
Jessica McAfee led a review describing the many different types of multiplex reporter assays and how they will be useful for finding causal genetic variants associated with brain traits. This work was in collaboration with the lab of Hyejung Won. The study was published in the Journal of Developmental Disorders.
Inferring imprinting during human neurogenesis
Dan Liang led a study inferring cell-type specific imprinting during human neurogenesis. This study was published at Human Molecular Genetics. These sites may be particularly useful to interpret parent of origin genetic association studies, as they start getting well powered.